Classification and Evolution:

The members of togaviridae are ssRNA (+) viruses that have spherical icosahedral shapes. Because of the tendency for this virus to be transmitted by insects, flaviviruses were originally mistakenly included under the umbrella of togaviridae. This original classification schema broke the virus into two groups: Alphaviruses and Flaviviruses; upon the advent of DNA sequencing, togaviruses were shown to have their replication (non-structural) proteins at their 5’ end, while their capsid proteins were localized to the 3’ end. At the present, there are two genera within the Togaviridae family: Alphavirus and Rubivirus. The Alphavirus genera consists of about 30 recognized members, whereas Rubivirus is home to only rubella.
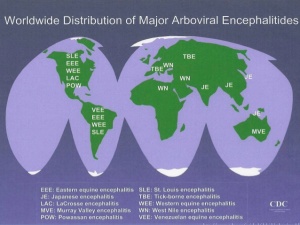
Alphaviruses have a worldwide distribution, including remote locations such as Antarctica; for this, Alphaviruses can be classically described as New World or Old World depending on their distribution. Most Alphaviruses are arboviral in nature, however, newly identified salmonid Alphaviruses such as salmon pancreas disease virus and sleeping disease virus are some examples of non arboviral transmission. Further sequencing has revealed that these viruses have diverged early on in Alphavirus evolution and have no present day close relatives. Another recent discovery of the southern elephant seal virus isolated from lice-infested seals have presented significant evidence for the marine origin of Alphaviruses.
Despite being classified in the same family, Rubiviruses and Alphaviruses have a clouded evolutionary relationship. While their genomes are similarly organized and capsids are physically homologous, they have radically different replication and assembly strategies.
Nomenclature:
Togaviruses are named after the drape-like way their envelopes cover their capsids.

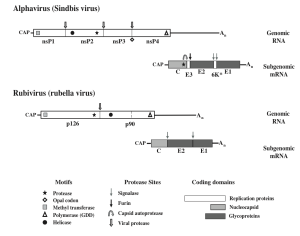
Structure:
The alphavirus capsid has been extensively studied with cryo-electron microscopy. Virions tend to be ~70nm in diameter and have a molecular mass of 52 Mega Daltons. Capsid composition is from repeating E1 and E2 glycoproteins, nucleocapsid protein C, genomic RNA, and a lipid envelope taken from the host. The proteins are arrange ad a T = 4 icosahedral lattice with 240 copies of each subunit, interacting together to form a rigid structure across the membrane. Also in the iral particle is a membrane association protein called 6K and another small protein called TF. The envelope stolen from the host is enriched with cholesterol and other molecules that facilitate with budding and entry.
The togavirus genome is a ssRNA (+) molecule that has a 5’ 7-methylguanisine cap and a 3’ polyadenated tail. Alphavirus genomes are about 2kb longer than their rubivirus counterparts. The first 2/3 of the genome at the 5’ end are dedicated to replication factors, whereas the final 1/3 at the 3’ end is reserved for structural proteins.
Replication:

Replication of Togaviruses is one of the major distinguishing factors from other viral species. Replication of alphaviruses occurs typically of RNA viruses: in the cytoplasm. After receptor-mediated endocytosis of the viral particle, the vesicle is transported to the lysosome, where the low pH catalyzes the uncoating of the genome, marking the beginning of the translational phase. The non-structural encoding 5’ end is translated into a polyprotein (P1234) which is hydrolytically cleaves into four distinct proteins (NSP1, a methylation and capping protein; NSP2, a helicase and protease; NSP3, a protein that converts cellular RNA replicase into plus stranded replicase; and NSP4, an RNA polymerase). These proteins work in conjunction to rapidly produce the complimentary RNA strand.
The RNA (-) strand produced by the replication proteins is used as the template for the production of the genomic RNA and subgenomic mRNA that contains the information for capsid production. This strategy allows for a great number of viral capsid proteins to be produced after an initial replication step.
The protein encoded by the mRNA is also a large polyprotein that is then cleaved into smaller components that come together and form the capsid. Capsid protein C is the first to be cleaved from the polyprotein; the remaining protein fragment is transported to the lumen of the ER, where host proteins process the remaining proteins.
New virus particles are assembled in the cytoplasm: genomic RNA contains an encapsulation signal, allowing it to selectively become surrounded in capsid proteins, forming a new virion. The viruses then escape the cell via budding.
Ribuviruses is similar in the sense that they produce subgenomic mRNA to handle capsid formation. Protein processing, however, is different in the rubivirus, as it relies on host signalase to cleave the capsid polyprotein. Structural glycoproteins are sequestered in Golgi body where a majority of viral assembly occurs.
Epidemiology:

Four alphaviruses have been responsible with human disease: Eastern Equine Encephalitis (EEE), Western Equine Encephalitis (WEE), Venezuelan Equine Encephalitis (VEE), and Chikungunya virus (CHIKV). These viruses have reservoirs in birds, birds and mammals, mammals, and primates, respectively. The Equine Encephalitis viruses cause flu-like symptoms, headaches, pain, and encephalitis, whereas CHIKV can cause rash, arthritis, fever, and headache.
Only one rubivirus is known to cause disease: Rubella. Rubella is extraordinarily contagious and is a known teratogen. Normally associated as a disease of childhood, Rubella can cause high fever, joint pain, headache, rash, and swelling if the lymph nodes.

Alphaviruses are transmitted primarily by mosquitos; Rubella is transmitted through airborne aerosolized droplets.
Rubella is one of the diseases covered by the MMR vaccine; while Rubella was at one point almost eradicated from the United States, the recent anti-vaccine movement has caused a significant resurgence. There are no current vaccines for alphaviruses.
Sources:
http://www.ncbi.nlm.nih.gov/books/NBK8200/
https://microbewiki.kenyon.edu/index.php/Togaviridae
http://www.virology.net/Big_Virology/BVRNAtoga.html
http://www.cdc.gov/vaccines/pubs/pinkbook/rubella.html
By Alex Lindqwister